Structural changes of tetrameric flavoprotein WrbA upon flavin binding
J. Wolfová1,2, J. Brynda1,3, J. R. Mesters4, J.
Carey5,
1Institute
of Physical Biology, University of South Bohemia České Budějovice, Zámek 136,
CZ-373 33 Nové Hrady, Czech Republic
2Institute of Systems Biology and Ecology, Academy of Science of the Czech Republic, Zámek 136,CZ-373 33 Nové Hrady, Czech Republic
3Institute of Molecular Genetics, Academy of
Sciences of the Czech Republic, Flemingovo nám. 2, CZ-16637
4Institute of Biochemistry, Center for Structural and Cell Biology in Medicine, University of Lübeck, Ratzeburger Allee 160, 23538 Lübeck, Germany
5Chemistry
Department,
julinka.w@tiscali.cz
Protein WrbA from Escherichia coli studied in this work represents a widely distributed family of tetrameric flavoenzymes [1, 2]. Using flavin mononucleotide (FMN) like monomeric flavodoxins that transfer single electrons to protein partners but forming multimers and carrying out two-electron reduction of quinones [3, 4] like the FAD-dependent quinone oxidoreductases, WrbA was suggested to be a structural and functional linker between bacterial flavodoxins and eukaryotic NAD(P)H:quinone oxidoreductases [5].
Interesting changes in protein dynamics and multimerization state accompanying FMN binding were identified by biophysical spectral methods [6]. This was motivation for the comparative analysis of the FMN-bound WrbA structure (holoWrbA) and the FMN-free WrbA structure (apoWrbA), results of which are presented here.
Structures of holoWrbA and apoWrbA were solved from x-ray diffraction data on single crystals and refined to a resolution of 2.0 Å and 1.85 Å, respectively. Although crystals for both protein forms were grown from the same crystallization solution, different space groups were determined: P41212 for holoWrbA and P21212 for apoWrbA.
In each structure four WrbA monomers form a tetramer, where individual subunits share the common fold of flavodoxins with sequence insertions unique for WrbA family forming additional secondary structure elements. FMN-binding sites are located at the interfaces of three of the four subunits. Our comparative analysis of holo- and apoWrbA revealed significant changes at the level of quaternary and tertiary structure. Large differences were observed in the arrangement of subunits in tetramers (see Fig. 1). In orthorhombic apoWrbA structure the distance between subunits across the empty FMN-binding site is larger by 2 to 4 Å than in holoWrbA structure, with the distances being longer at the surface than at the core of apoWrbA tetramer (see Fig. 2). Changes in structural organization of tetramers thought to be induced by FMN binding are in correspondence with the result of mass-spectrometry analysis [6] suggesting FMN to favour tetramer formation.
Substantial structural changes of WrbA monomers upon FMN binding are presumably located in the vicinity of the FMN-binding site. Structural overlay of holo- and apoWrbA monomers (see Fig. 3) shows the large relative motion of one of the loops contacting FMN in holoWrbA, resulting in partial occupation of the empty FMN-binding pocket in apoWrbA. FMN was also found to induce shifts in the positions of residues interacting with FMN, the most apparent being rotation of the side chain of Arg 78.
The 3D-superposition of holo- and apoWrbA with long-chain holo- and apoflavodoxin from Anabaena [7, 8] revealed striking similarities in the behavior of the FMN-binding residues in response to FMN binding, which are beyond those expectable from their distant homology. This finding indicates WrbA to be a significant member rather than a remote and unusual branch of flavodoxin-like proteins.
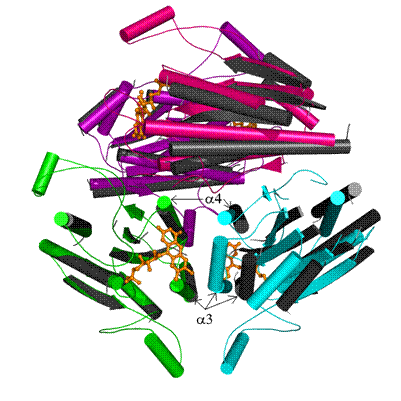
Figure 1. Overlay of holo- and apoWrbA tetramers. HoloWrbA
subunits are green, cyan, pink and purple, all apoWrbA subunits are dark grey. a helices are drawn as cylinders, b strands as arrows, loops as lines.
FMN cofactor in the holoWrbA structure is shown in ball and stick
representation and colored orange. Larger distances between subunits across the
empty FMN-binding site in apoWrbA tetramer bring the dark grey, cyan and pink
subunits out of alignment, while the green and purple subunits of holoWrbA are
aligned with the dark grey apoWrbA subunits nearly perfectly.
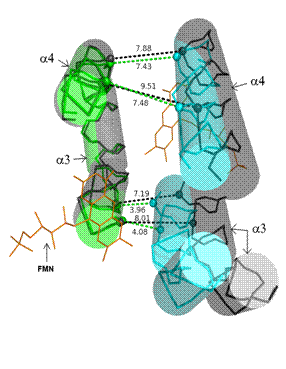
Figure 2. Interface between subunits formed across the
FMN-binding site in holo- and apoWrbA tetramers. The detailed picture was drawn out of the overlay
of WrbA tetramers shown in Fig. 1, thus colors and graphical representation are
the same. Polypeptide main-chain atoms included in a helices are shown in skeletal mode. Helices
denoted as a3 and a4, labeled also in Fig. 1, were
oriented to give an optimal view on the shift between subunits of apoWrbA
relative to holoWrbA. Distances between the corresponding main-chain atoms (atoms shown as spheres) of
the interfacing subunits are marked with dashed lines: green for holoWrbA, black
for apoWrbA; their lengths measured in Å indicate not only translational
but also rotational shift of the subunits upon FMN binding.
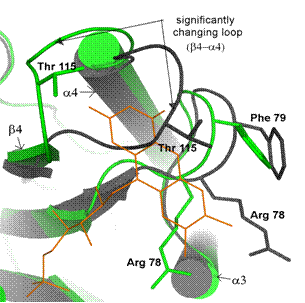
Figure
3. Overlay of holo- and apoWrbA
monomers – detail of FMN-binding site. Colors and graphical representation are
the same as in Fig. 1; holoWrba – green, apoWrbA – dark grey. Substantial
changes accompanying FMN binding are indicated: 1) loop between b4 and a4 occupying partly the empty FMN-binding site
in apoWrbA; 2) residues observed in different positions in holo- and apoWrbA are
labeled and their side chains are shown as skeletal models. Selected
secondary-structure elements are labeled.
All figures included in this paper were prepared by using the PyMOL molecular graphic system [9].
References
1. R. Grandori & J. Carey, Protein Sci., 3 (1994) 2185-2193.
2. R. Grandori, P. Khalifah, J.A. Boice, R.
Fairman, K. Giovanelli & J. Carey, J. Biol. Chem., 273 (1998)
20960-20966.
3. E.V. Patridge & J.G. Ferry, J.
Bacteriol., 188 (2006) 3498-3506.
4. G. Nöll, E. Kozma, R. Grandori, J. Carey, T. Schödl, G. Hauska & J.
Daub, Langmuir, 22 (2006) 2378-2383.
5. J. Carey, J. Brynda, J. Wolfova, R. Grandori, T. Gustavsson, R. Ettrich & I. Kuta Smatanova, Protein Sci., 16 (2006) 2301-2305.
6. A. Natalello, S.M. Doglia, J. Carey &
R. Grandori, Biochemistry, 46 (2007) 543-553.
7. S.T. Rao, F. Shaffie, C. Yu, K.A. Satyshur,
B.J. Stockman, J.L. Markley & M. Sundarlingham, Protein Sci., 1 (1992)
1413-1427.
8. C.G. Genzor, A. Perales-Alcon, J. Sancho
& A. Romero, Nat. Struct. Biol., 3 (1996) 329-332.
9. W.L. DeLano: The PyMOL Molecuular Graphic
System. San Carlos 2002. DeLano Scientific.
Acknowledgements.
This work was supported by the
Ministry of Education of the