GLYCOSYLATION
OF IgG-Fc
P. Kolenko1,2, J. Dohnálek1, J. Dušková1,
T. Skálová1, J. Hašek1
1Institute of Macromolecular
Chemistry AS CR, v.v.i., Heyrovského nám.2, 162 00, Prague 6
2Dept.of Solid State Physics, FJFI,
CTU, Trojanova 13, 120 00, Prague 2
kolenko@imc.cas.cz
Keywords: Fc, crystal, X-ray diffraction,
structure, saccharides
Introduction
The crystallisable fragment (Fc) of antibody (Ig) mediates the response
of the adaptive part of immune system. Ten structures of IgG-Fc in non-liganded
form were deposited in the Protein Data Bank [1] up to now. Although chemical
properties and the structure of Fc were assessed by many physical and chemical
techniques, some new details of the oligosaccharide structure were known after
evaluation of the most recently deposited structure [2] and our non-deposited
structure [3].
Localization of fucose
Interpretation of saccharides in electron density maps is difficult.
Inspection of structures and electron density maps showed doubtful structure
refinement of fucose. All structures of Fc deposited by December 2006 contained
beta-L-fucose. Low-resolution structural data did not allow distinguishing the
proper glycoform, e.g. two of the structures with electron density maps are
shown in Fig. 1.
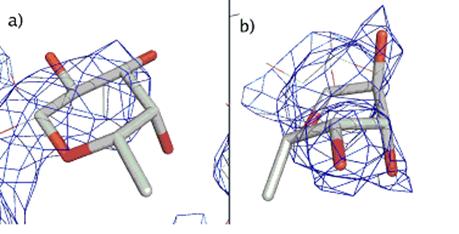
Figure 1. Electron density map in surroundings of fucose
for structures: a) 1I1C, b) 1H3U. The figures were prepared with Pymol [4].
In May 2007, the structure of non-liganded IgG1-Fc with the highest
resolution of 2 Å [2] was deposited in
the PDB. Our structure solution of IgG2b-Fc was described two years ago [3].
The structure contains alpha-L-fucose. The experimental data (high resolution
limit lower than 2.2 Å) allowed localization of fucose with good agreement with electron
density
(Fig.2).
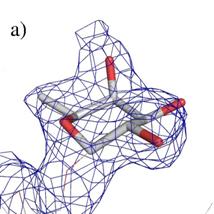
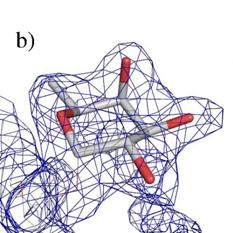
Figure 2. Electron density map in surroundings of fucose
for structures: a) 2DTQ, b) our structure. The electron density observed in the
lower left corner belongs to the continuation of the oligosaccharide chains in
both cases.
About 30 % of all PDB entries containing saccharides have at least one error
in glycan interpretation [5]. The glycosylation and quality of structures deposited
in the PDB will be discussed.
References
1. H.M. Berman, J. Westbrook, Z. Feng, G.
Gililand, T.N. Bhat, H. Weissing, I.N. Shindyalov, P.E. Bourne, Nucleic
Acids Research, 28, 2000, 235-242.
2. S. Matsumiya, Y. Yamaguchi, J. Saito, M.
Nagano, H. Sasakawa, S. Otaki, M. Satoh, K. Shitara, K. Kato, Journal of
Molecular Biology, 368, 2007, 767.
3. P.
Kolenko, J. Dohnálek, R. Šťouračová, T. Skálová, G. Tiščenko, J. Dušková, J.
Hašek, Materials Structure, 12, 2005, 146.
4. W.L. DeLano, The PYMOL User’s
Manual, DeLano Scientific, San Carlos, CA, USA, 2002.
5. T. Lütteke, M. Frank, C-W. von der Lieth, Carbohydrate Research, 339, 2004.
Acknowledgements
The
project is supported by the Ministry of Education, Youth and Sports of the
Czech Republic (project no. 1K05008) and by the project “Spine 2 – Complexes”
of the European Commission (LSHG-CT-2006-031220).