3DPatch: fast sequence and structure residue-level
information content annotation in a web browser
David Jakubec1,2,∗, Jiří Vondrášek1, Robert D. Finn3
1Department of
Bioinformatics, Institute of Organic Chemistry and Biochemistry of the Czech
Academy of Sciences, 166 10 Prague 6, Czech Republic
2Department of Physical
and Macromolecular Chemistry, Faculty of Science, Charles University,
128 43 Prague 2, Czech Republic
3European Molecular
Biology Laboratory, European Bioinformatics Institute (EMBL-EBI), Wellcome
Trust Genome Campus, Hinxton, Cambridge CB10 1SD, UK
∗To whom correspondence
should be addressed. Email: david.jakubec@uochb.cas.cz
Amino acid residues manifesting high levels
of conservation are often indicative of functionally
significant regions of protein structures.
Residues critical for protein folding, hydrophobic core stabilization,
intermolecular recognition, or enzymatic activity often manifest lower mutation
rates compared to the rest of the protein. Quantitative assessment of residue
conservation typically involves querying a sequence against a database, finding
similar sequences, aligning them to bring equivalent positions into register,
and applying an information theory-based measure to individual columns in the
multiple sequence alignment. Understanding how the sequence conservation
profile relates in 3D requires its projection onto a protein structure, which can
be a time-consuming process.
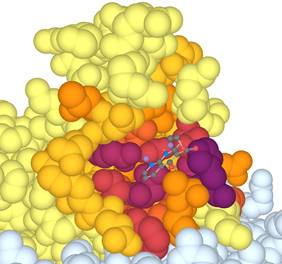
We developed 3DPatch, a client-side web
application that simplifies the task of calculating protein sequence
information content, 3D structure identification, and conservation level-based
mark-up (Figure 1). 3DPatch utilizes the power of profile hidden Markov models
and speed of HMMER3.1 to provide accurate results in a matter of seconds. It
was developed with easy integration into other peoples’ websites in mind and
supports most modern web browsers. 3DPatch is freely available at http://www.skylign.org/3DPatch/.