Structure of multifunctional plant nuclease TBN1
T. Kovaľ1, J. Stránský1,
2, T. Podzimek3, P. Lipovová3, J. Matoušek4, K.
Fejfarová1,
P. Kolenko1, J. Dušková1, T. Skálová1,
J. Hašek1 and J. Dohnálek1
1Institute of Macromolecular Chemistry, AS CR, v.v.i.,
Heyrovského nám. 2, 162 06 Praha 6,
2Faculty of Nuclear Sciences and Physical Engineering, Czech Technical University, Břehová 7, 115 19
Praha 1, Czech Republic
3Institute of Chemical Technology, Technická 5, 166 28 Praha 6, Czech Republic
4Institute of Plant Molecular Biology, Biology Centre, AS CR, v.v.i., Branišovská 31, 370 05 České Budějovice, Czech Republic
koval.tomas@gmail.com
Plant nuclease TBN1 (UniProt sequence accession no. Q0KFV0) from Solanum lycopersicum (tomato) is a Zn2+- dependent glycoprotein with a molecular mass of 31.6 kDa (about 37 kDa when glycosylated). TBN1 belongs to plant nuclease I family and plays an important role in specific apoptotic functions, vascular system development, stress response and tissue differentiation in plants [1]. In addition, TBN1 exhibits anticancerogenic properties [2].
Two recombinantly
expressed variants of TBN1 (wild type and hypoglycosylated
mutant N211D) were used in our study. Datasets for structural analysis were
collected at the synchrotron radiation source BESSY II (
TBN1 is mainly α-helical with a trinuclear Zn2+ cluster placed
in the active site in the center of the wide groove. Three oligosaccharides
bonded on the surface serve primarily as a shielding of the hydrophobic regions
and therefore contribute to solubility and stability of the enzyme (Figure
1). TBN1 acts as phosphodiesterase
cleaving the bond between phosphorus and
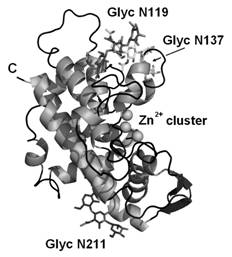
Figure 1 Fold, secondary structure
and main features of TBN1 [5].
The work on this project was
supported by the Czech Science Foundation, projects no. P302/11/0855,
202/06/0757 and 521/09/1214, by the EC under ELISA grant agreement number
226716 (synchrotron access, projects 09.2.90262 and 10.1.91347), by the Institution research plan AV0Z50510513 of the Institute
of Plant Molecular Biology, Biology Centre. We acknowledge support of the
Ministry of Education, Youth and Sports of the
1. J. Matousek, P. Kozlova, L. Orctova, A. Schmitz, K. Pesina, O. Bannach, N. Diermann, G. Steger,
D. Riesner, Biol. Chem., 388, (2007), 1–13.
2. J. Matousek, T. Podzimek, P. Pouckova, J. Stehlik, J. Skvor, P. Lipovova, J. Matousek, Neoplasma, 57,
(2010), 339-348.
3. T. Koval, P. Lipovova, T. Podzimek, J. Matousek, J. Duskova, T. Skalova, A. Stepankova, J. Hasek, J.
Dohnalek, Acta. Cryst. F67, (2011), 124-128.
4. J. Dohnalek, T. Koval, P. Lipovova, T. Podzimek, J. Matousek, J. Synch. Rad., 18, (2011), 29-30.
5. T. Koval, P. Lipovova, T. Podzimek, J. Matousek, J. Duskova, T. Skalova, A. Stepankova, J. Hasek, J.
Dohnalek, Acta. Cryst. D69, (2013), 213-226.